Full Length Research Article
Mitochondrial genetic characterization of Gujar population living in the Northwest areas of Pakistan
Inam Ullah*1, Habib Ahmad1,4, Brian E. Hemphill2, Muhammad Shahid Nadeem3, Muhammad Tariq4, Sadia Tabassum1
Adv. life sci., vol. 4, no. 3, pp. 84-91, May 2017
*– Corresponding Author: Dr. Inaam Ullah (Email: inamgenetics@gmail.com)
Authors' Affiliations
2- Department of Anthropology, University of Alaska, Fairbanks, Fairbanks, AK – United States
3- Department of Biochemistry, Faculty of Science, King Abdulaziz University Jeddah 21589 – Saudi Arabia
4- Islamia College University, Peshawar, Khyber Pakhtunkhwa – Pakistan
Abstract![]()
Introduction
Methods
Results
Discussion
References
Abstract
Background: Diversity of communities with specific cultural, ethnic, lingual and geographical backgrounds makes Pakistani society a suitable study subject to unravel the early human migrations, evolutionary history of population having about 18 ethnic groups. Gujars are mostly Indic-speaking nomadic herders with the claims of multiple origins in the sub-continent. Present study was aimed at the determination of maternal lineage of Gujars by mitochondrial DNA analysis.
Methods: Total DNA from the human buccal cells was isolated using modified phenol chloroform method. Purified DNA was used for the PCR amplification of mitochondrial Hyper Variable Region 1 and 2 (HVR1 & 2). The nucleotide sequences of amplified PCR products were used to explore the maternal lineage of the Gujar population residing in Northern Pakistan.
Results: Haplotypes, allele frequencies and population data of the mitochondrial control region was determined in 73 unrelated individuals belonging to Gujar ethnic group of Northwest areas of Pakistan. Total 46 diverse haplotypes were identified out of which 29 were found unique with (0.9223) genetic diversity and (0.9097) power of discrimination. Haplogroup R was the most frequent (48%) followed by haplogroup M (45%) and N (7%).
Conclusion: We found that the Gujar population has multiple maternal gene pool comprising of South Asian, West Eurasian, East Eurasian, Southeast Asian and fractions of Eastern Asian, Eastern Europe and Northern Asian lineages. This study will contribute for the development of mitochondrial DNA database for Pakistani population.
Key words: Pakistan, Swat, Gujar, mtDNA control region, Haplotyping
Introduction
Pakistan is located in the western part of the Indian subcontinent, with Afghanistan and Iran to the west, India to the east, the Arabian Sea to the south and covers an area of approximately 796,095 sq. km (figure 1). About 46,8000 sq. km of this area is in west and north comprises mountains lands and plateau, while the remaining 328,000 km2 is in the form of plains [1]. Pakistan has a diverse communities distributed into variety of ethnic groups, having variety of cultures, languages and geographical backgrounds, which make this land suitable for unraveling early human migrations, population study and evolutionary history having 18 ethnic groups further divided into casts and sub-casts [2, 3].
Gujars are Indic-speaking nomadic herders whose origins are claimed to be in Rajasthan and adjacent regions of Gujarat in India and the Indus Valley of Pakistan [4]. Following irrigation efforts in the Indus Valley by the British administration, Gujars were forced northwards in the late-19th century into the foothills rimming the northern margin of the Indus Valley and beyond into Khyber Pakhtunkhwa, Jammu and Kashmir. Some historians says that Gujars probably first appeared in the area about 400 years ago [5, 6]. Gujars are considered as ‘Aryas’ and their arrival to this part of the world is traced back to 242 and 300 BCs. Gujars invaded India in third century B.C. and they are actually inhabitants of Gujarustan which is still called as Gujarustan or Gorgia [7]. First time the word Gujar was used by a pioneer Ramchand with his name [8].
Various studies have proved that human DNA is a direction to explore historical movements of populations by studying their genetic make-up. Mitochondrial DNA is a proper tool for the human migration, geographic distribution and population origin due to its high evolutionary importance [9, 10].
To investigate all possible lineages among various ethnic groups, we obtained data for the Hyper Variable Region 1&2 (HVR1&2) of mtDNA from 73 Gujar individuals from the Swat district of Khyber Pakhtunkhwa Pakistan. mtDNA haplogroups affiliations have been diagnosed by using different computer software and servers and finally we compared the mtDNA distribution among the various subpopulations, including regional ethnic groups from Pakistan and neighboring countries.
Methods
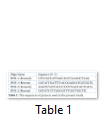
Saliva samples were collected in sterile collecting cups from 73 unrelated Gujar volunteers belongs to different areas of district Swat of Northwest Pakistan (figure 1). All participants gave their informed consent verbally or in writing after explaining the aims and procedures of the study to them. The consent form was designed according to the ethical review board of Hazara University. Genomic DNA from the human buccal cells was obtained using DNA isolation method [11]. The isolated genomic DNA was used for the PCR amplification of HVR1 & 2 of mtDNA with two sets of reverse and forward primers (table 1). The PCR reaction mixture included 2.0µL of 10pM/µL F-Primer, 2.0µL of 10pM/µL R-Primer, 0.5µL of Taq DNA Polymerase enzyme (5U/µL) “Fermentas”, and 2.0µL of DNA template with a final volume of 25.0µL. Thermal cycling was conducted using an Applied Bio system 2720 (95°C for 4 min; 35 cycles of 94°C for 40 s, 56°C for 1 minute, and 72°C for 1 minutes; and a final extension at 72°C for 5 min). The gel containing PCR products were purified using the procedure adopted from GeneAll Gel Elution Kit (SV) Cat. no. 102-101. Sequencer machine (ABI Prism 3730XL) was used for sequencing the purified products.
Data analysis
Haplotypes for the corresponding HVR1 and HVR2 sequences were then identified with the help of online software, MitoTool [12], HaploGrep [13] and Mitomaster [14] using PhyloTree Build 16 (http://www.phylotree.org) as classification tree to assess the quality of mtDNA data [10]. The sequences of Gujar mitochondrial DNA were assign to haplogroup according to phylotree [10] and published data [15-18]. The population statistics i.e. Genetic Diversity (GD), Power of Discrimination (PD) and Random Match Probability (RMP) were also calculated using computational tools [19, 20].
Results
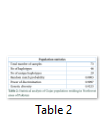
A total of 73 samples were analyzed for the mitochondrial DNA control region of Gujar population belongs to District Swat of Khyber Pakhtunkhwa (KP) Province of Pakistan. Haplogroup frequencies were calculated for the characterization of mtDNA variation in the individuals of the present study population. Forty six different haplotypes were observed during the present study among which 29 were unique while 17 haplotypes were shared by more than one individual, while the corresponding mtDNA genetic diversity was (0.9223), power of discrimination (0.9097) and random match probability (0.0903) table 2. The observed haplogroup frequencies, their respective variants and geographic position are given in table 3.
By comparing the genetic parameters of the reported population living in Pakistan with the current studied Gujar population, we found that the Gujars of Swat have a moderate unique haplotypes (29) consistent with the other population of Pakistan (table 4). The moderate frequency of unique haplotypes reflected in high genetic diversity (0.922) in the Gujar ethnic group of the present study as compared to the other reported ethnic groups from Pakistan except Kalash with (0.851) genetic diversity (table 4). However, the highest number of unique haplotypes (128) has also been reported in Pakhtuns of Pakistan due to large number of sample size (n= 230) table 4.
The obtained sequences of mtDNA control region (1-574, 15974-16425) of the present Gujar population were compared with revised Cambridge Reference Sequence (rCRS) [21]. The results of sequences revealed that at nucleotide position 16023np 95% (G/A), at 16061np 91% (C/A), at 16163np 95% (G/A) , at 32np 92% (A/G), at 38np 98.5% (G/A) and at 278np 100% (A/G) had transition mutations while transversion mutations were scored at 16036np 99% (G/C), 16172np 100% (G/T), 16219np 97% (A/G), 33np 95% (C/G), 44np 93% (C/A) respectively.
In the present study we observed South Asian haplogroups (42%), West Eurasian (37%), East Eurasian (11%), Southeast Asian (4%), Eastern Asian (2.7%), Eastern Europe (1.4%) and Northern Asian (1.4%). Among south Asian haplogroups, haplogroup M6 occurred (7%), M30 (4%), M37 (4%), M5c (4%), M3 (2.7%), M3a (2.7%), M5 (2.7%), M52a (2.7%), R5a (2.7%), M30d (1.4%), M3c (1.4%), M53 (1.4%), M54 (1.4%), M7c (1.4%) and R22 (1.4%). West Eurasian haplogroups includes H2a (4%), T2b (4%), H14a (2.7%), H5 (2.7%), K1a (2.7%), U7a (2.7%), H1 (1.4%), H1a (1.4%), H1e (1.4%), H3p (1.4%), N (1.4%), T (1.4%), T1a (1.4%), U2a (1.4%), U4a (1.4%), U5b (1.4%), U7 (1.4%), V9a (1.4%) and W3a (1.4%). East Eurasian haplogroups includes B4a (5%), D4b (1.4%), D4e (1.4%), D4g (1.4%) and D4p (1.4%). Southeast Asian haplogroups includes F1 (1.4%), G2b (1.4%) and S (1.4%). Eastern Asian haplogroups includes A (2.7%); Eastern Europe H7i (1.4%) and Northern Asian include haplogroup J (1.4%) respectively. The frequencies of each haplogroups are given in (figure 2).
The haplotypes of Gujar population were assigned to mega haplogroups which revealed that the most frequent among them was R with the frequency of (48%) followed by haplogroup M (45%) and N (7%) (figure 3).
Tables & Figures
Discussion
In the present study 73 unrelated samples from the Gujars were characterized for maternal linage and other genetic structure. The genetic structure of the present studied population was compared with the previously reported data of Pakistani ethnic groups. The haplotypic diversity of the Gujar population (GD=0.9223) observed shows a high genetic diversity in comparison with the other reported population of Pakistan except Kalash [22-25]. Genetic diversity is due the reflection of unique haplotypes distribution. The numbers of unique haplotypes identified in the present studied population were 63%, which were found somehow consistent with Burusho 78%, Hazara 76%, Makrani 76%, Baluchi 69% and Brahui 68% among the other reported population of Pakistan, while moderately lower from Saraiki 92%, Sindhi 90% and Pathan 81% [22-25]. Members of Gujars population revealed high frequency (42%) of South Asian lineage. The proportion of South Asian lineages in the other reported Pakistani populations were 48% in Sindhi, 39.1% in Pathan, 36% Pashtun, 29.4% in Saraiki and 24% in Makrani [22, 24-27]. Low frequency of South Asian lineages among the major ethnic groups of Afghanistan have also been reported with the prevalence of 15% in Hazara, 13.3% in Baluch and 7.1% in Pashtun, while absent in Tajik [28]. The presence of south Asian mtDNA haplogroups in the present study population revealed that the population residing in this region are the true inhabitants and are remolded in the past by local demographic events [17]. The West Eurasian haplogroup was the second most prevalent haplogroup accounting for (37%) in the individuals of the present study population. Its frequency among the Pathans of Pakistan was reported 55%and 26% in Makranis [24, 25]. Furthermore, the frequency of West Eurasian haplogroup in Indian Punjabis population were reported from (40-50%), in Kashmiris and Gujrathis 30%, while the least were observed in Indian Uttar Pradesh and West Bengal [17,29]. Greater proportion of West Eurasian lineages were also reported among the major ethnic groups of Afghanistan with the frequencies of 40% in Hazara, 89% in Tajik, 74% in Baluch and 64% in Pashtun [28] . The presence of these lineages revealed that, the gene flow in the past to this region may occur from the west through Iran or from the North through Central Asia [23], through the invasion by different invaders i.e. Alexander, Arabians, Muslims and the British [30]. The mega haplogroup R, M and N identified in the Gujars population are said to be South Asian in origin and has been originated approximately 60000-75000 years ago in South Asia [31], suggesting their maternal gene pool as South Asian in origin.
Acknowledgment
The authors would like thank the Ethnogenetic Project (No. 20-1409) titled “Ethnogenetic elaboration of KP through dental morphology and DNA analysis” at Hazara University, Mansehra, Pakistan for assisting in sample collection. The research was funded by the Indigenous 5000 Ph.D. Fellowship Program of the Higher Education Commission of Pakistan.
References
- Ahmad K, Hussain M, Ashraf M, Luqman M, Ashraf MY, et al. Indigenous vegetation of Soone Valley; At the risk of extinction. Pak J Bot, (2007); 39(3): 679-690.
- Ayub Q, Tyler-Smith C. Genetic variation in South Asia: assessing the influences of geography, language and ethnicity for understanding history and disease risk. Briefings in functional genomics & proteomics, (2009); 8(5): 395-404.
- Grimes BF. Ethnologue: languages of the world: 1992. Dallas, Texas: Summer Institute of Linguistics. Inc
- Grierson GA. 1903–1928. Linguistic Survey of India, Vols I-XI.: 1968. Calcutta[Reprint 1968, Delhi: Motilal Benarsidass]
- Barth F. Ecologic relationships of ethnic groups in Swat, North Pakistan. American Anthropologist, (1956); 58(6): 1079-1089.
- Rome S. Forestry in the princely state of Swat and Kalam (North-West Pakistan). 2005: pp. 1-125.
- Ali I. Mapping and documentation of the cultural assets of Kaghan Valley, Mansehra. United Nations Educational, Scientific and Cultural Organization, Islamabad, (2005); 5-6.
- Chauhan RAH. A short history of the Gurjars: past and present/by Rana Ali Hasan Chauhan. (2001).
- Nesheva D. Aspects of ancient mitochondrial DNA analysis in different populations for understanding human evolution. science, (2014); 1(5): 5-14.
- Van Oven M, Kayser M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Human mutation, (2009); 30(2): E386-E394.
- Akbar N, Ahmad H, Nadeem MS, Ali N, Saadiq M. An Efficient Procedure for DNA Isolation and Profiling of the Hyper Variable MtDNA Sequences. Journal of Life Sciences, (2015); 9530-534.
- Fan L, Yao Y-G. MitoTool: a web server for the analysis and retrieval of human mitochondrial DNA sequence variations. Mitochondrion, (2011); 11(2): 351-356.
- Kloss‐Brandstätter A, Pacher D, Schönherr S, Weissensteiner H, Binna R, et al. HaploGrep: a fast and reliable algorithm for automatic classification of mitochondrial DNA haplogroups. Human mutation, (2011); 32(1): 25-32.
- Brandon MC, Ruiz‐Pesini E, Mishmar D, Procaccio V, Lott MT, et al. MITOMASTER: a bioinformatics tool for the analysis of mitochondrial DNA sequences. Human mutation, (2009); 30(1): 1-6.
- Behar DM, Villems R, Soodyall H, Blue-Smith J, Pereira L, et al. The dawn of human matrilineal diversity. The American Journal of Human Genetics, (2008); 82(5): 1130-1140.
- Elmadawy MA, Nagai A, Gomaa GM, Hegazy HM, Shaaban FE, et al. Investigation of mtDNA control region sequences in an Egyptian population sample. Legal Medicine, (2013); 15(6): 338-341.
- Metspalu M, Kivisild T, Metspalu E, Parik J, Hudjashov G, et al. Most of the extant mtDNA boundaries in south and southwest Asia were likely shaped during the initial settlement of Eurasia by anatomically modern humans. BMC genetics, (2004); 5(1): 26.
- Oven M, Vermeulen M, Kayser M. Multiplex genotyping system for efficient inference of matrilineal genetic ancestry with continental resolution. Investigative Genetics, (2011); 2(6): 1-14.
- Prieto L, Zimmermann B, Goios A, Rodriguez-Monge A, Paneto G, et al. The GHEP–EMPOP collaboration on mtDNA population data—A new resource for forensic casework. Forensic Science International: Genetics, (2011); 5(2): 146-151.
- Tajima F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics, (1989); 123(3): 585-595.
- Andrews RM, Kubacka I, Chinnery PF, Lightowlers RN, Turnbull DM, et al. Reanalysis and revision of the Cambridge reference sequence for human mitochondrial DNA. Nature genetics, (1999); 23(2): 147-147.
- Hayat S, Akhtar T, Siddiqi MH, Rakha A, Haider N, et al. Mitochondrial DNA control region sequences study in Saraiki population from Pakistan. Legal Medicine, (2015); 17(2): 140-144.
- Quintana-Murci L, Chaix R, Wells RS, Behar DM, Sayar H, et al. Where west meets east: the complex mtDNA landscape of the southwest and Central Asian corridor. The American Journal of Human Genetics, (2004); 74(5): 827-845.
- Rakha A, Shin K-J, Yoon JA, Kim NY, Siddique MH, et al. Forensic and genetic characterization of mtDNA from Pathans of Pakistan. International journal of legal medicine, (2011); 125(6): 841-848.
- Siddiqi MH, Akhtar T, Rakha A, Abbas G, Ali A, et al. Genetic characterization of the Makrani people of Pakistan from mitochondrial DNA control-region data. Legal Medicine, (2015); 17(2): 134-139.
- Bhatti S, Aslamkhan M, Abbas S, Attimonelli M, Aydin HH, et al. Genetic analysis of mitochondrial DNA control region variations in four tribes of Khyber Pakhtunkhwa, Pakistan. Mitochondrial DNA Part A, (2016); 1-11.
- Bhatti S, Aslamkhan M, Attimonelli M, Abbas S, Aydin HH. Mitochondrial DNA variation in the Sindh population of Pakistan. Australian Journal of Forensic Sciences, (2017); 49(2): 201-216.
- Whale J. Mitochondrial DNA analysis of four ethnic groups of Afghanistan. (2012). University of Portsmouth.
- Ahmed M. Ancient Pakistan-an archaeological history. 2014 Amazon.
- McElreavey K, Quintana-Murci L. A population genetics perspective of the Indus Valley through uniparentally-inherited markers. Annals of human biology, (2005); 32(2): 154-162.
- Kivisild T, Rootsi S, Metspalu M, Mastana S, Kaldma K, et al. The genetic heritage of the earliest settlers persists both in Indian tribal and caste populations. The American Journal of Human Genetics, (2003); 72(2): 313-332.